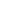
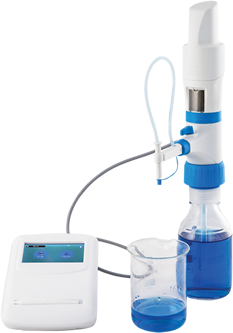
RNA-sequencing (RNA-seq) is a scientific assay that is used to quantify and examine RNA in a biological sample using next-generation sequencing techniques. The process analyzes the transcriptome to identify the state of genes in our DNA, whether they are turned on or off, as well as the degree of gene activation. Let’s take a look at the working principle of this technique, along with its various applications and challenges.
Working Principle of RNA Sequencing
At the outset, RNA sequencing was performed using Sanger sequencing technology, which is not a cost-and time effective technique in current times. Recently, with the advent of next-generation sequencing (NGS) technology, this assay can be performed at low costs without compromising on the overall throughput.
There are several steps involved in an RNA-seq workflow, including:
- RNA extraction
- Reverse transcription into cDNA
- Adapted ligation
- Amplification
- Sequencing
1) After the extraction of RNA from a sample, the first step is the conversion of RNA of interest into complementary DNA (cDNA) fragments to form a cDNA library. This step is performed through reverse transcription.
2) This is followed by fragmentation of cDNA, and addition of adapters to these fragments. The adapters consist of functional elements that enable the process of sequencing. For instance, the priming and amplification element. After this step, processes, such as amplification, size selection and clean-up are performed.
3) Then, the cDNA library is examined using NGS, which produces short sequences corresponding to the respective fragment.
The depth of RNA sequencing can vary according to the output data requirements. Sequencing can be performed by either single-end or paired-end sequencing techniques. Single-read sequencing is cheaper and faster as the cDNA fragments are sequenced from only one of the ends. In the paired-end technique, sequencing is performed from both ends, but can prove to be beneficial in post-sequencing data reconstruction.
RNA-seq versus Microarrays
RNA-seq is a highly sophisticated technique and is considered to be relatively better than other technologies, including microarray hybridization. The possible reasons for the superiority of RNA-sequencing over other techniques, include:
1) RNA sequencing is not limited by previously identified genomic sequences. Hybridization-based approaches usually require species specific probes, while RNA sequencing is able to identify novel transcripts from entities which have not been sequenced before.
2) The isolated cDNA sequences can be mapped directly to the specific regions in the genome, making the process less erroneous by eliminating the background signal.
3) In microarray experiments, there can be problems related with cross-hybridization or sub-standard hybridization, which are not a major concern in RNA sequencing experiments.
4) Unlike microarray data results, which are relative values calculated with respect to background signals, RNA sequencing data is absolute and quantifiable.
5) Microarrays are ineffective for detection of extremely high or low transcription levels, which is not an issue with RNA sequencing.
Applications of RNA Sequencing
RNA sequencing is useful in examining and discovering various aspects of the transcriptome, such as the mRNA, rRNA and tRNA content. Unraveling the transcriptome can help us co-relate the information stored in the genome with the functional protein expression. In addition, it has the ability to reveal what genes are activated in the genome, as well as their degree of transcription. These discoveries enable researchers to comprehend a cell’s biology in a better way and identify factors that lead to disease. The most common techniques that use RNA sequencing include single nucleotide polymorphism (SNP) identification RNA editing, differential gene expression analysis and transcriptional profiling.
Challenges and Future Perspectives
Although there have been significant strides in the field of RNA sequencing in the last few years, there are various challenges that remain unaddressed.
RNA sequencing requires isolation of sufficient and high-quality RNA, which leads to disposal of a significant portion of a sample. While the sample quantity requirements have minimized substantially over the years, one needs to ensure that they have isolated sufficient amounts of RNA that can fulfill all their data analysis needs. Poor quality samples usually give unsatisfactory outcomes.
Further, RNA degrades rapidly, therefore, it is vital to handle RNA samples with utmost caution at every step of the process. In order to evaluate a sample’s quality and concentration, UV-visible spectroscopy techniques are used.
In an effort to reduce sequencing costs and enable sequencing of small samples, scientists pool samples before the library preparation step. However, while analyzing data, researchers need to account for this factor, and if there are variations within the pooled samples, it can generate misleading results and cause statistical issues.
Over the decade, there has been significant advancement in the domain of RNA sequencing, which is validated through increased throughput and reduced costs. Currently, the sequence fidelity in the process is much better than earlier available NGS technologies. This is accompanied by wider availability of data analysis tools. However, sequencing technologies still need to be improved in order to obtain sufficient read depth and maximize sequencing capacity to increase the number of biological replicates that can be evaluated to generate meaningful data.




 11018
11018